Climbing performance is a great measure of Drosophila (fruit fly) health. It requires the coordination of many muscular, neural, developmental, and physiological systems, making it a strong metric of health used for organismal-level phenotyping in genetic, pharmacological, and aging studies.

FreeClimber
Researchers often use manual techniques to measure either the average height climbed over a period of time or the time required for a fraction of all flies to reach a given height. While analyzing videos this way is simple, it is both tedious and time-consuming. Computational approaches seek to automate this analytic process, but most published methods are written in a proprietary language (ex. MATLAB), have limited availability beyond a lab group, or necessitate costly and special filming setups. Each drawback is a hurdle to the broader deployment of computational approaches.
As part of my dissertation, I developed FreeClimber [link]: a novel, automated, and high-throughput climbing analysis pipeline. Unlike previous other methods, FreeClimber is an open-source and publicly available program built with Python and does not require a custom setup. This program introduces several new features that also improve on previous methods:
- Implementation: Equipped to run as either a Graphical User Interface (GUI) for individual video processing and parameter optimization, and a Command Line Interface (CUI) for processing similar videos with common configuration settings.
- Pre-process: Improves data quality in videos with noisy environments, via background subtraction of static pixels.
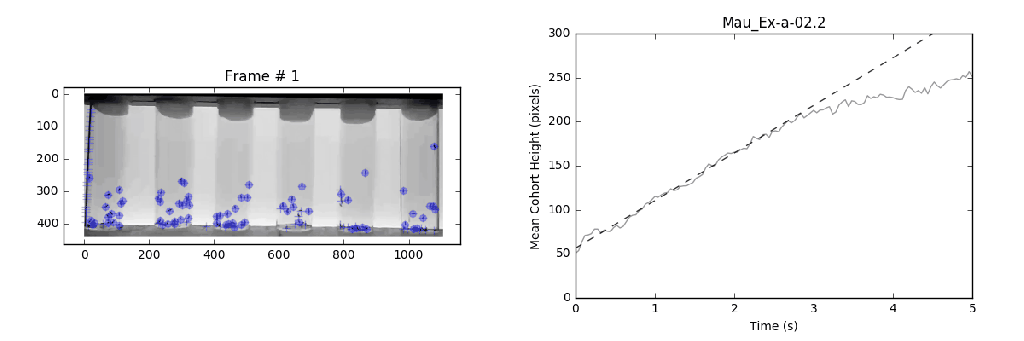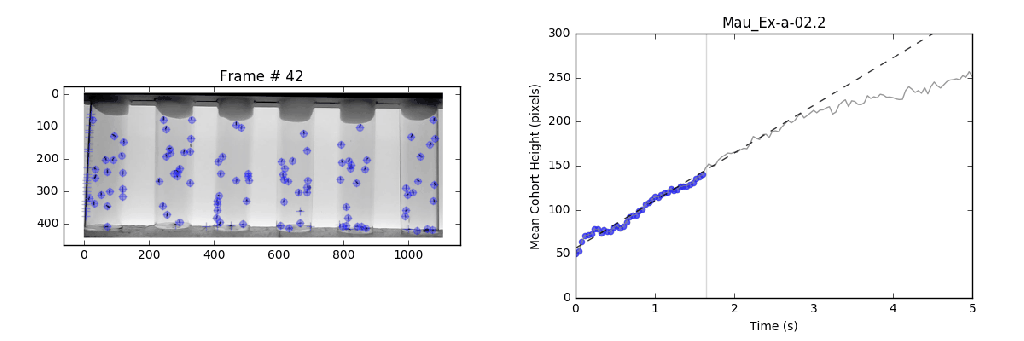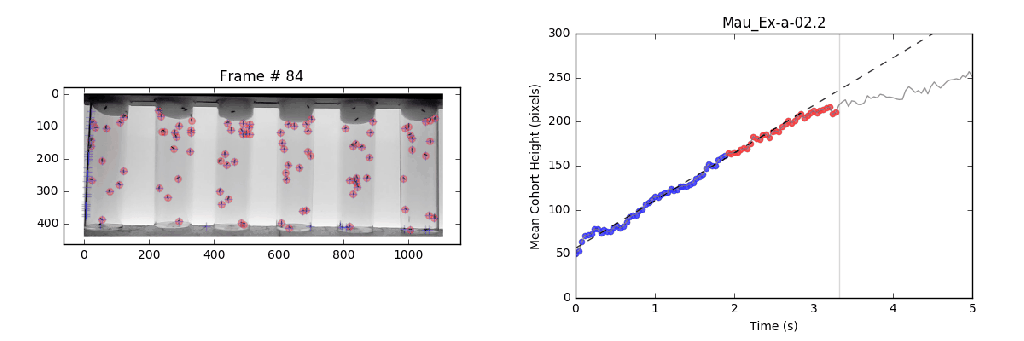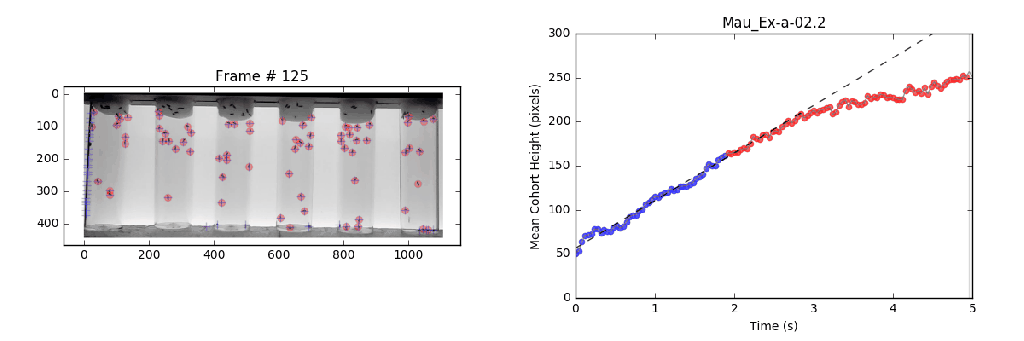
- Analysis: Circumvents previously violated assumptions from manual methods that lead to systematic bias in analysis with a local linear regression. This approach identifies the slope (velocity) for the most linear portion (greatest consistency) of a mean-vertical position vs. time curve.
- Post-process: Enables follow up with individual spot coordinates so more advanced analyses can be deployed with trackpy (tracking individual flies) or manual analyses.
These benefits all sum up to make FreeClimber more efficient and effective, and generate more repeatable, reproducible, and biologically-meaningful results than traditional manual methods of analysis.
Beyond my dissertation, FreeClimber was the recipient of the Brown University 2020 Library Innovation for Innovations in Research Rigor, Transparency or Reproducibility in Life and Physical Sciences [link]. This work is available through the Journal of Experimental Biology [link] and is currently in use by several labs investigating lipid metabolism, neurodegenerative diseases, and more.
Studying higher-order genetic interactions
Prior to the development of FreeClimber, I helped establish a collaboration between the Rand lab (mitochondrial-nuclear genetic interactions) and the Wessells lab at Wayne State University (genetics of exercise conditioning) [link]. Together, we sought to understand how mitochondrial-nuclear genome interactions impacted flies’ ability to respond to a longitudinal exercise conditioning program, and how various molecular and physiological biomarkers corresponded with differences in climbing performance.
Results from this study are described in our 2019 Mitochondrion paper “Mito-nuclear interactions modify Drosophila exercise performance” [link], which I can pass along upon request [link].