In addition to my dissertation work, I was also part of a National Science Foundation’s Integrate Graduate Education Research Traineeship (NSF-IGERT). This traineeship sought to bring scientists from biology, geology, and mathematics together to answer highly interdisciplinary, student-driven questions related to Reverse Ecology [link]. This field aims to connect genomic variation with functional phenotypic variation in a natural setting.
Genomic sequencing and transcriptomic analysis of parasitized vs. unparasitized amphipods
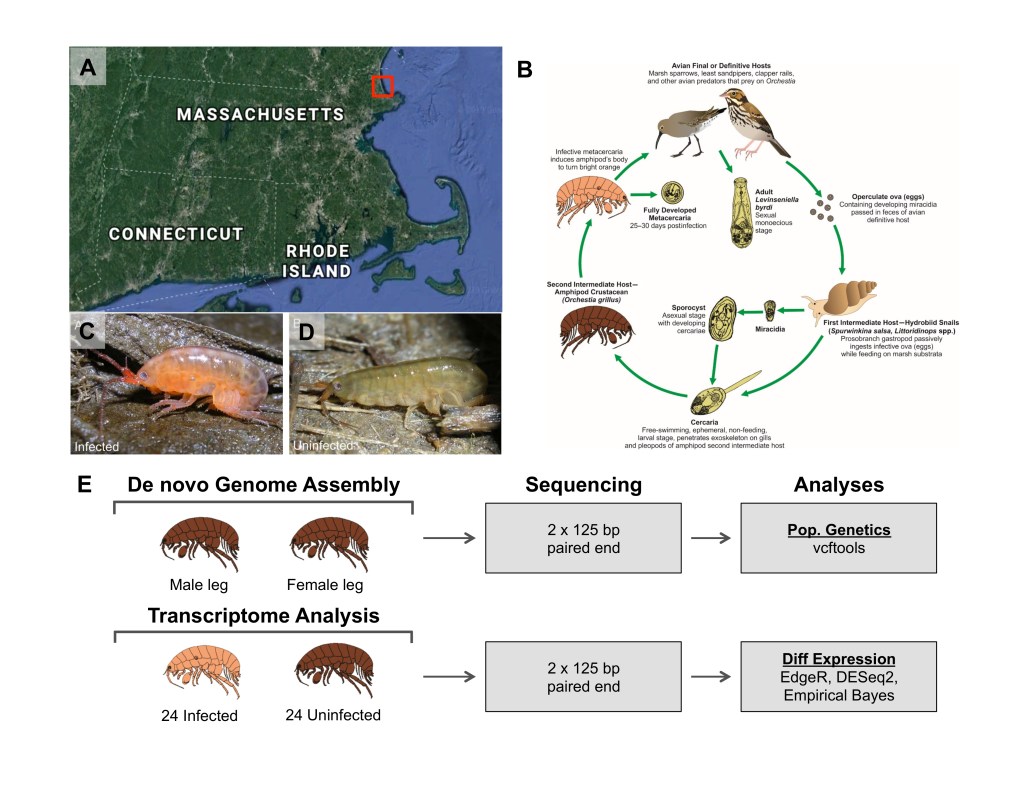
My Fellowship cohort investigated a mind controlling host-parasite interaction between an amphipod (Orchestia grillus) and a parasitic trematode (Levinseniella byrdii) [link] at the Plum Island Estuary Long Term Ecosystem [link]. Normally, amphipods hide under leaves and eat detritus (decaying organic matter). But when they are infected by L. byrdii, these amphipods become an intermediate host in the parasite’s life cycle. As the parasite matures, the amphipod’s camouflaged patterning becomes a bright orange-red and the amphipod loses its aversion to light. They are also more likely to climb tall stalks of marsh grass and wave their forelegs, drawing the attention of their predator and the parasite’s final host: the sandpiper [link].

Uninfected amphipod with three different helminths for scale. (Left) Amphipods measure 2-3 cm in length and can serve as a host to several different helminth parasites. Three of these parasites are pictured in their cyst stage relative to the host. (Right) Three helminth species include Levinseniella byrdii (top), an unidentified species of Microphallus (middle), and an unidentified species of Metameritrema (bottom). Amphipods infected with L. byrdii are typically red in color, like the one pictured in the figure below.
Our group was intrigued by this system and set out to sequence the amphipod genome and identify the transcriptomic differences between infected and uninfected individuals. We identified significant changes in gene expression for genes known to affect immune-response, phototaxis (light-directed locomotion), and oxidative phosphorylation complexes (energy production).
This work is published in Molecular Ecology [link].
Uncovering metagenomic signatures from a salt pond sediment core
I was also fortunate to help the following year’s cohort in their project looking at the metagenomic signature of a meter-long sediment core from a New England salt pond [link]. The group sought to understand how biogeochemical changes corresponded with differences in microbiome community assembly in a coastal salt pond. My focus sought to determine if there was a connection between microbial diversity and hurricane disturbance based on historical records and geological evidence.