Insect flight is one of the most energetically demanding phenotypes in the animal kingdom and is important for foraging, mate searching, predator evasion, migration, etc. Flight is also a complex trait, meaning it is the result of multiple different genetic elements working together. Identifying the genetic modifiers of flight is especially challenging because of the shear number of elements with small to moderate effects. Most phenotypes and genetic disorders in humans are also complex and have important genetic/cellular/molecular crossover with Drosophila. These similarities make the fly a highly relevant model for understanding the biological basis of complex traits and diseases.
Across two chapters of my dissertation, I sought to identify the genetic modifiers for insect flight ability and variability. To measure flight performance, I used a flight column to abruptly drop groups of flies into an adhesive-lined cylinder [link]. Flies react to the sudden drop by righting themselves and flying to the cylinder edge. Stronger flies react sooner, respond more effectively, and land higher than weaker flies. The corresponding mean landing height for all flies in a sex-genotype combination was deemed that group’s flight ability, while the coefficient of variation (mean normalized-standard deviation) was deemed the group’s flight variability (more technically referred to as robustness of flight).

I tested flight performance across nearly 200 unique genotypes in the Drosophila Genetic Reference Panel [link] and performed a Genome Wide Association Study (GWAS). I also used several approaches to map additive and epistatic (genes interacting with other genes) variants, whole genes, and gene-gene interaction networks.
I identified over a thousand independent genetic markers associated development for both flight ability and variability. Namely, I found associations for transcription factors (genes regulating the expression of other genes), genes involved in general development (wings, muscles, sensory hair structures), and neurodevelopment of the sensory and peripheral nervous systems. One of my most significant and novel findings was an under-appreciated role for smell and taste receptor genes in flight-based proprioception and the wiring of complex and sex-specific neural circuits in modifying flight performance.
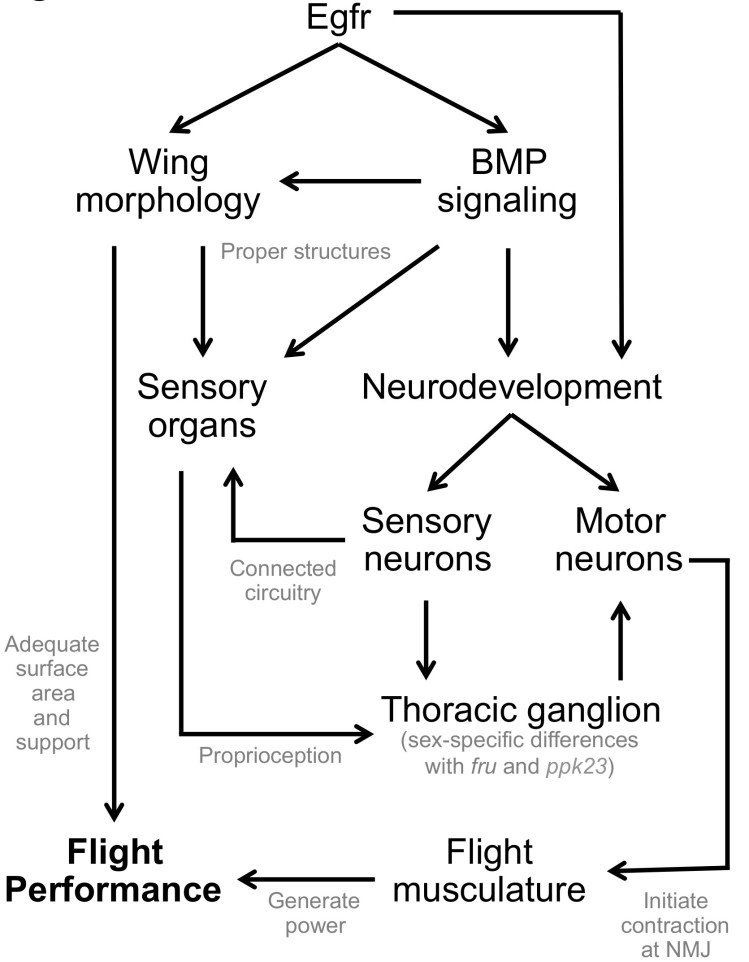
Working model for genetic modifiers affecting flight ability in a natural population. Fight performance has a complex genetic architecture, with the key developmental gene Epidermal Growth Factor Receptor (Egfr; human homolog EGFR) and the BMP signaling pathway contributing to wing and neurodevelopment. Both are both important for structuring the sensory organs that enable the fly to use mechanosensory channels for proprioception. Signals from the sensory organs on the wing, head, and body travel to the brain and thoracic ganglion. This ganglion acts as a flight control center, sending other signals through the motor neurons to the direct and indirect flight musculature for control of the wing angle and generation of power during flight, respectively. Functional genetic variation across genotypes was detected at each of these structures in our study.
Abbreviations: BMP = bone morphogenetic protein; fru = fruitless (neurodevelopmental gene involved in courtship); ppk23 = pickpocket 23 (human homolog ASIC5, involved in chemosensation of pheromones and mechanosensation/proprioception); NMJ = neuromuscular junction (interface between neuron and muscle).
In developing my working model, I co-opted several inferential approaches previously used only in human and human cell culture studies. One of these methods was PEGASUS [link], which assesses whole gene significance from a distribution of individual genetic variants’ significance scores. I retooled PEGASUS to work with Drosophila inputs, and validated one gene in the lab and found previous studies validated two others. We released our novel application of PEGASUS as PEGASUS_flies, currently available on the Ramachandran lab Github [link].
Both works are currently available through PLOS Genetics [flight performance (ability) ] or are in review and available through the bioRxiv preprint server [robustness of flight (variability)].